Many years ago, the US Naval Ocean Systems Center (NOSC) in San Diego, CA, was in need of a better way to access data. The University of Hawaii had assembled a very large collection of marine biological data in the Hawaiian archipelago. The collection, known as the Hawaiian Coastal Zone Data Bank (HCZDB), contained 65 million individual observations of marine life found in the waters surrounding Hawaii. The amount of data overpowered all existing methods of analysis. The data was available but could not be accessed efficiently enough to perform detailed analyses.
One of my employer's west coast cost centers decided to
fund an Independent Research and Development (IR&D)
project to solve NOSC's
problem. The center director knew me from my work on the
east coast. At the time, I had been programming for two
years in the production environment (with five years prior
military programming in COBOL, RPG, and Auto Coder). I was
considered an expert in the COBOL and FORTRAN programming
languages. But what probably caught the director's eye
was my work in winning a major military contract
DPSC
 .
.
Upon his request, and with the concurrence of a few key
employees, I moved to the west coast. The project became
named the Biological Data Bank (BIODAB). I assembled a
programming team that was charged with providing a
solution. It seemed that everyone had read
E. F. Codd's
 work and wanted the team to build a relational database
system. No such animal existed. Anywhere.
work and wanted the team to build a relational database
system. No such animal existed. Anywhere.
As development lead, I carved out the positions of architect and database crafter for myself.
One team member became the user interface (UI). There
were about 75 stakeholders, worldwide, all of whom had
suggestions as to how the database system should work. The
availability of email on the
ARPANET
 allowed these users to contact team members directly. The
bombardment of emails resulted in a loss of productivity.
We established that the UI
was the only team member who would answer user questions.
At architectural meetings, the UI
would provide the team with the users' suggestions,
most of which were adopted. An unexpected benefit of
assigning the user interface to one individual was that
most questions could be answered immediately (the
UI
was fully aware of questions asked earlier). This improved
the user's view of the development team (warranted or not).
allowed these users to contact team members directly. The
bombardment of emails resulted in a loss of productivity.
We established that the UI
was the only team member who would answer user questions.
At architectural meetings, the UI
would provide the team with the users' suggestions,
most of which were adopted. An unexpected benefit of
assigning the user interface to one individual was that
most questions could be answered immediately (the
UI
was fully aware of questions asked earlier). This improved
the user's view of the development team (warranted or not).
Another team member was assigned the responsibility of building the low level database access functions. The task was complicated by the fact that the actual database structure was in flux as I stitched the database together. The access functions were assembly routines that made the target machine's masked search instructions available to the higher level languages (FORTRAN and Pascal) as well as functions that would implement the search and extraction function.
The architecture took form during team architectural meetings. Basic modules such as searching, reporting, and extracting were supplemented by ancillary modules such as development schedule, "what's new", and help. There were no load or unload modules as these features would be incorporated in a follow-on (hopefully customer funded) contract. Although management disagreed, my approach to developing modules was to complete those that produced the biggest bang for the smallest buck. Thus, "what's new" was developed before searching. Developing somewhat simpler modules before very much more complex modules gave the team an opportunity to gain experience with the operating system in a simpler environment. This, in turn, improved team confidence and morale as more and more modules were implemented. It also gave our clients more and more working parts so that the user interface became second nature. The help facility proved its value many times over. It also gave me time to craft the database before a database was actually needed.
Since there were no tools to load the database, I had to develop them. The first task was to scrub all 65 million HCZDB records. The criteria were simple: a valid latitude, longitude, and depth triplet must be present; a valid date-time must be present; and the marine life that was observed must be able to be categorized using a biological taxonomy. The first criteria reduced the data to about 100 thousand records; the second reduced the data to about 65 thousand records. However, NOSC required that invalid date-time tagging be treated as missing data.
The actual database implementation was elusive. There
were a large number of alternatives from which to choose.
My task was helped by a serendipitous reading of
E. W. Dijkstra's monogram
Notes on Structured Programming
 .
At one point in the text, Dijkstra discussed how little
we, as programmers, know about the hardware that we
program and how much we must trust the hardware engineers.
His example was a 27-bit multiplier. As a thought
experiment, he suggested testing the multiplier by
supplying each possible input value as the multiplier and
each possible input value as the multiplicand. Assume that
each multiplication takes tens of microseconds to perform
(in those days, very fast) and that the result of the
multiplication could be validated instantly. It would take
almost ten thousand years to complete the test. Dijkstra
suggested that we, as programmers, take a good deal on
faith. As an aside, Dijkstra also suggested that we really
don't multiply that many unique numbers.
.
At one point in the text, Dijkstra discussed how little
we, as programmers, know about the hardware that we
program and how much we must trust the hardware engineers.
His example was a 27-bit multiplier. As a thought
experiment, he suggested testing the multiplier by
supplying each possible input value as the multiplier and
each possible input value as the multiplicand. Assume that
each multiplication takes tens of microseconds to perform
(in those days, very fast) and that the result of the
multiplication could be validated instantly. It would take
almost ten thousand years to complete the test. Dijkstra
suggested that we, as programmers, take a good deal on
faith. As an aside, Dijkstra also suggested that we really
don't multiply that many unique numbers.
That last conjecture hit me hard. Extending the concept would imply that we do not use very many unique values in a database. At first, I did not believe it! As a practical test, I accessed a database containing ocean column data (e.g., bathymetry, salinity, dissolved phosphates, etc.) at various latitudes and longitudes just off the Pacific coast. Ignoring the data units (i.e., feet, percentage, mg/L, etc.), I extracted 35 thousand values. When the values were normalized (by converting integers to real) and duplicates eliminated, about 2,500 unique values remained. Dijkstra's surmise had been substantiated. Now the question became "how can we apply this finding to a database design?"
After some thought, I concluded that data could be stored as surrogate values in a database. The surrogate value, usually significantly smaller than the value it represents, is actually an index into an attribute table.
For example, assume that a last name may contain up to 50 characters (allowing for hyphenated last names). If there are one million last names then this collection requires 50Mb of storage (1M last names at 50 bytes per).
Now assume that three-quarters of the last names are duplicates, yielding 250,000 unique values. The largest surrogate value for the collection would be 250,000. The binary value (111101000010010000) for this largest surrogate requires 18 bits. Rounding upward, we reserve three 8-bit bytes for each last name surrogate in the database. The surrogate values can be stored in the database using 3Mb (1M last names at three bytes per). To determine the surrogate value for a given last name, an attribute surrogate value table is required. For the last names attribute, this table requires 13.25Mb (12.5Mb for the unique last names and 750Kb for the surrogate value associated with each last name). All told, this example database requires 16.25Mb. This is a savings of about 65% (over the earlier 50Mb). In practice, the database normally compressed its raw data to 10% of its original size.
One of the reasons that the surrogate value database worked so well was because the Unisys 1100 instruction set included masked search instructions1:
| Masked Search Equal | Masked Search Not Equal |
| Masked Search Less than or Equal | Masked Search Greater Than |
| Masked Search for Within Range | Masked Search for Not Within Range |
Each of these instructions can compare up to 256K words (specified in a count register) for equality, non-equality, etc. with the contents of a register (specified in the instruction) after the register and the data have been ANDed with the contents of the mask register. The words do not have to be consecutive, but have to be at some constant stride. Note that the masked search instructions (with a few others for preloading constants, etc.) could check for a pattern match over a large amount of data in one instruction. Each instruction stops at the first match.
This last point was of interest. To most efficiently query a database for some attributes, separated by ANDs, the search should initially seek the value that occurs least often in the database. To this end, the table containing the attribute surrogate values also contained a count of the number of times that the attribute value appeared in the database. Consider the search
last_name == 'foster' and division == 'manufacturing'
Assume that the database structure is:
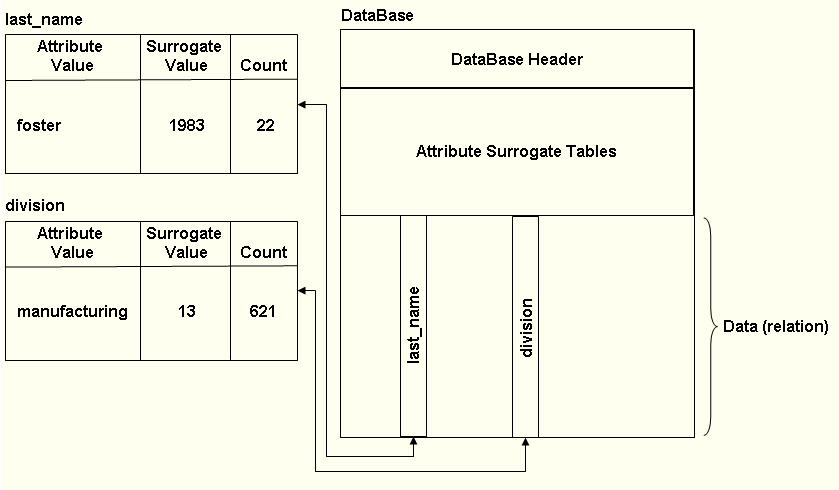
A masked search would first be executed for
"last_name == 'foster'"
When a match was found, a "found_counter" would be incremented, and the division attribute surrogate value in that record would be tested against the 'manufacturing' surrogate value. If, after processing the record, the "found_counter" value equaled the count in the last_name attribute surrogate value table for 'foster', the search would terminate; otherwise, the search would continue. Note that this search would have to stop and test the surrogate value for the division 22 times. Had the search first executed
"division == 'manufacturing'"
the search would have to stop and test the surrogate value in the last_name 621 times, at most failing 599 times. That search effort could be reduced if a count of the last_name hits were maintained. Then the search could terminate when the 22 instances of 'foster' were found.
Without a doubt, BIODAB was a success. Although it did not allow a user to load new data or unload existing data, BIODAB provided a fast, easy to use, relational database management system. It was successful enough that the NOSC decided to fund the follow-on project (the reason my employer funded the IR&D project in the first place).
BIODAB
also provided the team members with experience in
multi-language development. It also caused me to rethink
my views on what development language should be used in
other projects. My language of choice (Pascal) was
unfortunately destroyed by a very poor choice made by the
X3J9 Pascal technical committee
 .
.
The team member who was assigned the responsibility of building the low level database access functions was Tim Johnson. He recalls the BIODAB project and shared the following with me.
There are three aspects of this project that stand out for me more than others. First, was the cohesion of the team. We really had solid technical people who worked well together. The other two points are memories of some technical issues I helped solve.
The first was using the bank switching technique to reduce the memory footprint. If I remember right, you put me on to that, and it might have gone back as far as the Command, Control, and Communications project, but I know there was a major change for BIODAB due I think to a change in architecture between the 1108 and the 1100 series.
The other technical break-through that I've never forgotten is coming up with the algorithm for selecting the "isolated" AND attribute candidates in a complex predicate. If you remember, we were allowing search predicates that used parenthetically aggregated ANDs, ORs, and NOTs. We did this based on an algorithm that Ben Consilvio found for us for converting arithmetic predicates to Polish notation. The morning shower Aha that I had was that if the operands in the predicate were all set to false, except for one, and evaluated, and if the computed result was true, that operand (i.e., attribute) must be an isolated AND candidate. By looking at the counts you mention above, we picked the one that would have the fewest stops in the masked search.
1As an aside, there is interesting speculation as to why the Masked Search instructions came into existence on the Univac machines. There are many who suggest that National Security Agency was responsible.